Logistic Regression Assumptions and Diagnostics in R
The logistic regression model makes several assumptions about the data.
This chapter describes the major assumptions and provides practical guide, in R, to check whether these assumptions hold true for your data, which is essential to build a good model.
Make sure you have read the logistic regression essentials in Chapter @ref(logistic-regression).
Contents:
Logistic regression assumptions
The logistic regression method assumes that:
- The outcome is a binary or dichotomous variable like yes vs no, positive vs negative, 1 vs 0.
- There is a linear relationship between the logit of the outcome and each predictor variables. Recall that the logit function is
logit(p) = log(p/(1-p)), where p is the probabilities of the outcome (see Chapter @ref(logistic-regression)). - There is no influential values (extreme values or outliers) in the continuous predictors
- There is no high intercorrelations (i.e. multicollinearity) among the predictors.
To improve the accuracy of your model, you should make sure that these assumptions hold true for your data. In the following sections, we’ll describe how to diagnostic potential problems in the data.
Loading required R packages
tidyversefor easy data manipulation and visualizationbroom: creates a tidy data frame from statistical test results
library(tidyverse)
library(broom)
theme_set(theme_classic())Building a logistic regression model
We start by computing an example of logistic regression model using the PimaIndiansDiabetes2 [mlbench package], introduced in Chapter @ref(classification-in-r), for predicting the probability of diabetes test positivity based on clinical variables.
# Load the data
data("PimaIndiansDiabetes2", package = "mlbench")
PimaIndiansDiabetes2 <- na.omit(PimaIndiansDiabetes2)
# Fit the logistic regression model
model <- glm(diabetes ~., data = PimaIndiansDiabetes2,
family = binomial)
# Predict the probability (p) of diabete positivity
probabilities <- predict(model, type = "response")
predicted.classes <- ifelse(probabilities > 0.5, "pos", "neg")
head(predicted.classes)## 4 5 7 9 14 15
## "neg" "pos" "neg" "pos" "pos" "pos"Logistic regression diagnostics
Linearity assumption
Here, we’ll check the linear relationship between continuous predictor variables and the logit of the outcome. This can be done by visually inspecting the scatter plot between each predictor and the logit values.
- Remove qualitative variables from the original data frame and bind the logit values to the data:
# Select only numeric predictors
mydata <- PimaIndiansDiabetes2 %>%
dplyr::select_if(is.numeric)
predictors <- colnames(mydata)
# Bind the logit and tidying the data for plot
mydata <- mydata %>%
mutate(logit = log(probabilities/(1-probabilities))) %>%
gather(key = "predictors", value = "predictor.value", -logit)- Create the scatter plots:
ggplot(mydata, aes(logit, predictor.value))+
geom_point(size = 0.5, alpha = 0.5) +
geom_smooth(method = "loess") +
theme_bw() +
facet_wrap(~predictors, scales = "free_y")
The smoothed scatter plots show that variables glucose, mass, pregnant, pressure and triceps are all quite linearly associated with the diabetes outcome in logit scale.
The variable age and pedigree is not linear and might need some transformations. If the scatter plot shows non-linearity, you need other methods to build the model such as including 2 or 3-power terms, fractional polynomials and spline function (Chapter @ref(polynomial-and-spline-regression)).
Influential values
Influential values are extreme individual data points that can alter the quality of the logistic regression model.
The most extreme values in the data can be examined by visualizing the Cook’s distance values. Here we label the top 3 largest values:
plot(model, which = 4, id.n = 3)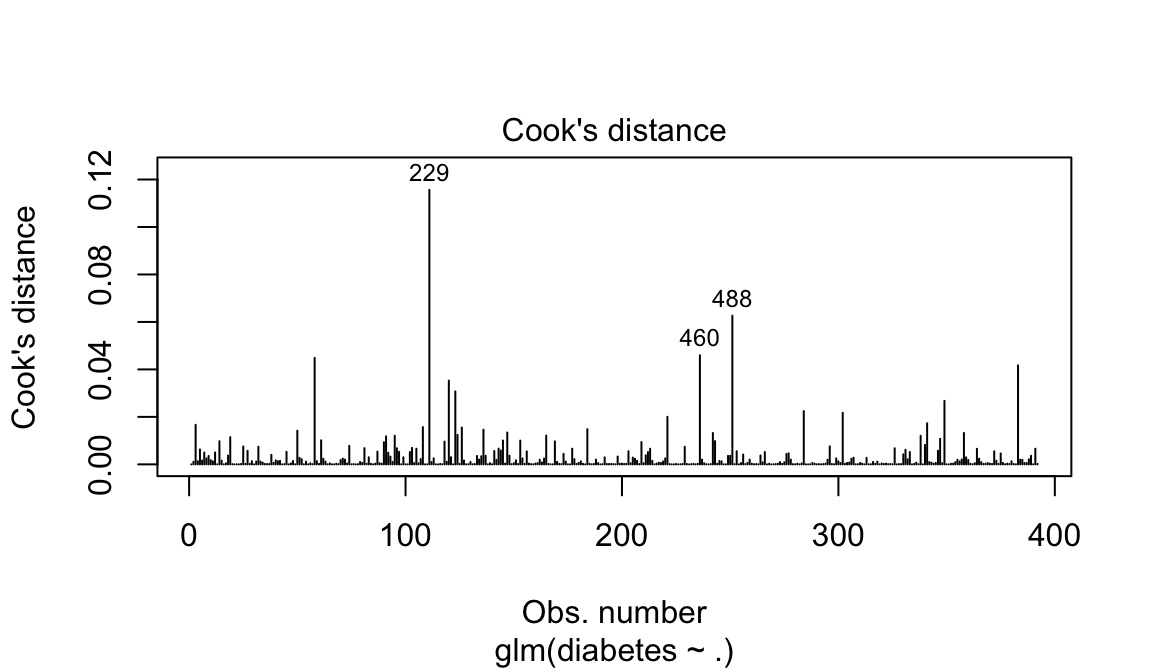
Note that, not all outliers are influential observations. To check whether the data contains potential influential observations, the standardized residual error can be inspected. Data points with an absolute standardized residuals above 3 represent possible outliers and may deserve closer attention.
The following R code computes the standardized residuals (.std.resid) and the Cook’s distance (.cooksd) using the R function augment() [broom package].
# Extract model results
model.data <- augment(model) %>%
mutate(index = 1:n()) The data for the top 3 largest values, according to the Cook’s distance, can be displayed as follow:
model.data %>% top_n(3, .cooksd)Plot the standardized residuals:
ggplot(model.data, aes(index, .std.resid)) +
geom_point(aes(color = diabetes), alpha = .5) +
theme_bw()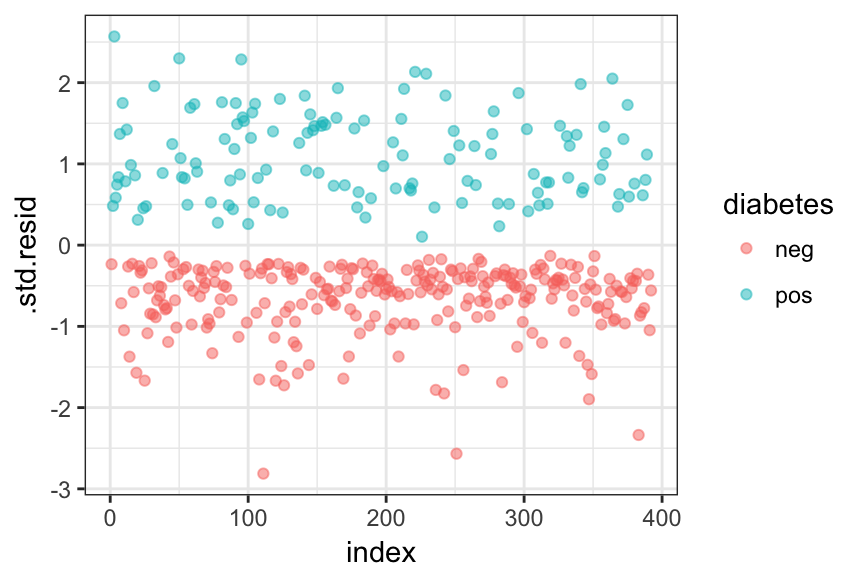
Filter potential influential data points with abs(.std.res) > 3:
model.data %>%
filter(abs(.std.resid) > 3)There is no influential observations in our data.
When you have outliers in a continuous predictor, potential solutions include:
- Removing the concerned records
- Transform the data into log scale
- Use non parametric methods
Multicollinearity
Multicollinearity corresponds to a situation where the data contain highly correlated predictor variables. Read more in Chapter @ref(multicollinearity).
Multicollinearity is an important issue in regression analysis and should be fixed by removing the concerned variables. It can be assessed using the R function vif() [car package], which computes the variance inflation factors:
car::vif(model)## pregnant glucose pressure triceps insulin mass pedigree age
## 1.89 1.38 1.19 1.64 1.38 1.83 1.03 1.97As a rule of thumb, a VIF value that exceeds 5 or 10 indicates a problematic amount of collinearity. In our example, there is no collinearity: all variables have a value of VIF well below 5.
Discussion
This chapter describes the main assumptions of logistic regression model and provides examples of R code to diagnostic potential problems in the data, including non linearity between the predictor variables and the logit of the outcome, the presence of influential observations in the data and multicollinearity among predictors.
Fixing these potential problems might improve considerably the goodness of the model. See also, additional performance metrics to check the validity of your model are described in the Chapter @ref(classification-model-evaluation).



