Principal component analysis : the basics you should read - R software and data mining
- What is principal component analysis?
- Main purpose of PCA
- Basic statistics - Covariance between two variables
- Covariance/correlation matrix
- Interpretention of the covariance matrix
- How to minimize the distortion in the data ?
- PCA terminologies : Eigenvalues / eigenvectors
- Steps for principal component analysis
- Compute principal component analysis (step by step)
- Packages in R for the principal component analysis
- Infos
What is principal component analysis?
Principal component analysis (PCA) is used to summarize the information in a data set described by multiple variables.
Note that, the information in a data is the total variation it contains.
PCA reduces the dimensionality of data containing a large set of variables. This is achieved by transforming the initial variables into a new small set of variables without loosing the most important information in the original data set.
These new variables corresponds to a linear combination of the originals and are called principal components.
This article describes, step by step, how PCA works using R software.
PCA basics
Understanding the details of PCA requires knowledge of linear algebra. In this section, we’ll explain the basics with simple graphical representation of the data.
In the Figure 1A below, the data are represented in the X-Y coordinate system. The dimension reduction is achieved by identifying the principal directions, called principal components, in which the data varies.
PCA assumes that the directions with the largest variances are the most “important” (i.e, the most principal).
In the figure below, the PC1 axis is the first principal direction along which the samples show the largest variation. The PC2 axis is the second most important direction and it is orthogonal to the PC1 axis.
The dimensionality of our two-dimensional data can be reduced to a single dimension by projecting each sample onto the first principal component (Figure 1B)

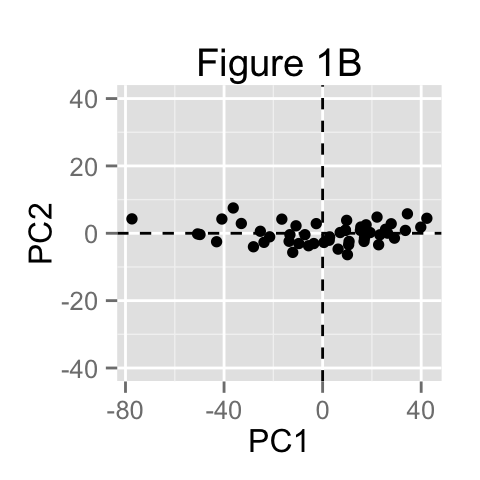
Main purpose of PCA
The main goals of principal component analysis is :
- to identify hidden pattern in a data set
- to reduce the dimensionnality of the data by removing the noise and redundancy in the data
- to identify correlated variables
PCA method is particularly useful when the variables within the data set are highly correlated.
Correlation indicates that there is redundancy in the data. Due to this redundancy, PCA can be used to reduce the original variables into a smaller number of new variables ( = principal components) explaining most of the variance in the original variables.
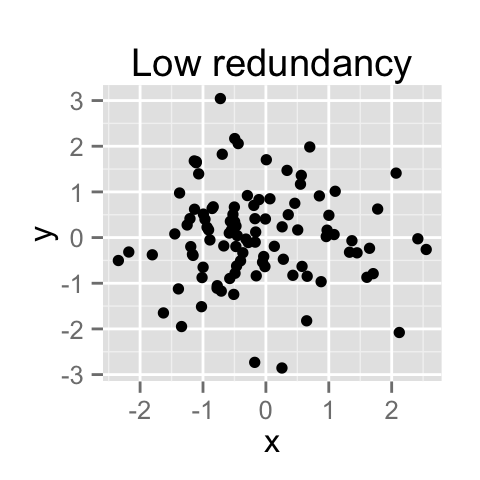
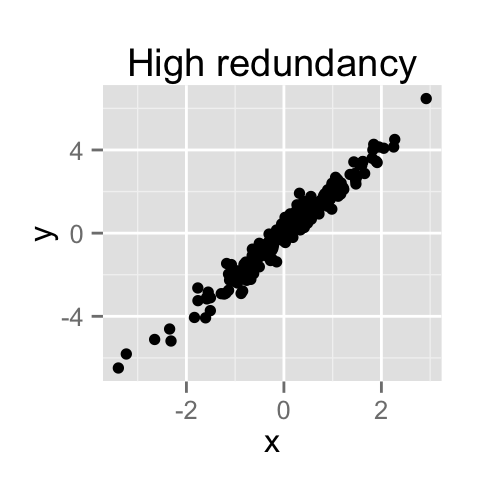
How to remove the redundancy?
PCA is traditionally performed on covariance matrix or correlation matrix.
Basic statistics - Covariance between two variables
Let x and y be two variables with length n.
The variance of x is :
\[\sigma^2_{xx} = \frac{\sum_i(x_i - m_x)(x_i - m_x)}{n - 1}\]
The variance of y is :
\[\sigma^2_{yy} = \frac{\sum_i(y_i - m_y)(y_i - m_y)}{n - 1}\]
The covariance of x and y is :
\[\sigma^2_{xy} = \frac{\sum_i(x_i - m_x)(y_i - m_y)}{n - 1}\]
\(m_x\) and \(m_y\) are the means of x and y variables, respectively.
The covariance measures the degree of the relationship between x and y.
Covariance/correlation matrix
A covariance matrix contains the covariances between all possible pairs of variables in the data set :
df <- iris[, -5]
res.cov <- cov(df)
round(res.cov,2) Sepal.Length Sepal.Width Petal.Length Petal.Width
Sepal.Length 0.69 -0.04 1.27 0.52
Sepal.Width -0.04 0.19 -0.33 -0.12
Petal.Length 1.27 -0.33 3.12 1.30
Petal.Width 0.52 -0.12 1.30 0.58Note that, the covariance matrix is symmetric. In the table above, covariance between Sepal.Length and Sepal.Width = covariance between Sepal.Width and Sepal.Length.
Interpretention of the covariance matrix
- The diagonal elements are the variances of the different variables. A large diagonal values correspond to strong signal.
diag(res.cov)Sepal.Length Sepal.Width Petal.Length Petal.Width
0.6856935 0.1899794 3.1162779 0.5810063 - The off-diagonal values are the covariances between variables. They reflect distortions in the data (noise, redundancy, …). Large off-diagonal values correspond to high distortions in our data.
The aim of PCA is to minimize this distortions and to summarize the essential information in the data
How to minimize the distortion in the data ?
In the covariance table above, the off-diagonal values are different from zero. This indicates the presence of redundancy in the data. In other words, there is a certain amount of correlation between variables.
This kind of matrix, with non-zero off-diagonal values, is called “non-diagonal” matrix.
We need to redefine our initial variables (x, y, z, ….) in order to diagonalize the covariance matrix.
This means that we want to change the covariance matrix so that the off–diagonal elements are close to zero (i.e, zero correlation between pairs of distinct variables).
The new variables (x’, y’, z’, …) are a linear combination of the old ones :
\[X' = a_1X + a_2Y + a_3Z, ...\]
\[Y' = b_1X + b_2Y + b_3Z, ...\]
In PCA, the constants a1, a2, an, b1, b2, bn are calculated such that the covariance matrix is diagonal.
PCA terminologies : Eigenvalues / eigenvectors
Eigenvalues : The numbers on the diagonal of the diagonalized covariance matrix are called eigenvalues of the covariance matrix. Large eigenvalues correspond to large variances.
Eigenvectors : The directions of the new rotated axes are called the eigenvectors of the covariance matrix.
Eigenvalues and eigenvectors can be easily calculated in R as follow :
eigen(res.cov)$values
[1] 4.22824171 0.24267075 0.07820950 0.02383509
$vectors
[,1] [,2] [,3] [,4]
[1,] 0.36138659 -0.65658877 -0.58202985 0.3154872
[2,] -0.08452251 -0.73016143 0.59791083 -0.3197231
[3,] 0.85667061 0.17337266 0.07623608 -0.4798390
[4,] 0.35828920 0.07548102 0.54583143 0.7536574The first principal components of the data are the first directions explaining maximum variances. This is equivalent to the first eigenvectors of the covariance matrix.
Steps for principal component analysis
The procedure includes 5 simple steps :
- Prepare the data :
- Center the data : subtract the mean from each variables. This produces a data set whose mean is zero.
- Scale the data : If the variances of the variables in your data are significantly different, it’s a good idea to scale the data to unit variance. This is achieved by dividing each variables by its standard deviation.
- Calculate the covariance/correlation matrix
- Calculate the eigenvectors and the eigenvalues of the covariance matrix
- Choose principal components : eigenvectors are ordered by eigenvalues from the highest to the lowest. The number of chosen eigenvectors will be the number of dimensions of the new data set. eigenvectors = (eig_1, eig_2,…, eig_n)
- compute the new dataset :
- transpose eigeinvectors : rows are eigenvectors
- transpose the adjusted data (rows are variables and columns are individuals)
- new.data = eigenvectors.transposed X adjustedData.transposed
Compute principal component analysis (step by step)
The data set iris is used : columns are variables and rows are observations:
df <- iris[, -5]
head(df) Sepal.Length Sepal.Width Petal.Length Petal.Width
1 5.1 3.5 1.4 0.2
2 4.9 3.0 1.4 0.2
3 4.7 3.2 1.3 0.2
4 4.6 3.1 1.5 0.2
5 5.0 3.6 1.4 0.2
6 5.4 3.9 1.7 0.41. Center and scale the data
df.scaled <- scale(df, center = TRUE, scale = TRUE)2. Compute the correlation matrix :
# 1. Correlation matrix
res.cor <- cor(df.scaled)
round(res.cor, 2) Sepal.Length Sepal.Width Petal.Length Petal.Width
Sepal.Length 1.00 -0.12 0.87 0.82
Sepal.Width -0.12 1.00 -0.43 -0.37
Petal.Length 0.87 -0.43 1.00 0.96
Petal.Width 0.82 -0.37 0.96 1.003. Calculate the eigenvectors/eigenvalues of the correlation matrix :
# 2. Calculate eigenvectors/eigenvalues
res.eig <- eigen(res.cor)
res.eig$values
[1] 2.91849782 0.91403047 0.14675688 0.02071484
$vectors
[,1] [,2] [,3] [,4]
[1,] 0.5210659 -0.37741762 0.7195664 0.2612863
[2,] -0.2693474 -0.92329566 -0.2443818 -0.1235096
[3,] 0.5804131 -0.02449161 -0.1421264 -0.8014492
[4,] 0.5648565 -0.06694199 -0.6342727 0.5235971The first eigenvalue (2.9) is much larger than the second (0.9), and so on…. The highest eigenvalues correspond to the first data principal components.
5. compute the new dataset :
# Transpose eigeinvectors
eigenvectors.t <- t(res.eig$vectors)
# Transpose the adjusted data
df.scaled.t <- t(df.scaled)
# The new dataset
df.new <- eigenvectors.t %*% df.scaled.t
# Transpose new data ad rename columns
df.new <- t(df.new)
colnames(df.new) <- c("PC1", "PC2", "PC3", "PC4")
head(df.new) PC1 PC2 PC3 PC4
[1,] -2.257141 -0.4784238 0.12727962 0.024087508
[2,] -2.074013 0.6718827 0.23382552 0.102662845
[3,] -2.356335 0.3407664 -0.04405390 0.028282305
[4,] -2.291707 0.5953999 -0.09098530 -0.065735340
[5,] -2.381863 -0.6446757 -0.01568565 -0.035802870
[6,] -2.068701 -1.4842053 -0.02687825 0.006586116Packages in R for the principal component analysis
There are several functions from different packages for performing PCA :
- The functions prcomp()and princomp() from the built-in R stats package
- PCA() from FactoMineR package. Read more here : PCA with FactoMineR
- dudi.pca() from ade4 package. Read more here : PCA with ade4
Infos
This analysis has been performed using R software (ver. 3.1.2) and ggplot2 (ver. 1.0.0)
Read more :
- Gregory B. Anderson, principal component analysis in R, https://www.ime.usp.br/~pavan/pdf/MAE0330-PCA-R-2013
- Wikibooks, http://en.wikibooks.org/wiki/Data_Mining_Algorithms_In_R/Dimensionality_Reduction/Principal_Component_Analysis
- Carlos Pinto, Data reduction, https://medicine.tcd.ie/neuropsychiatric-genetics/assets/pdf/2009_7_PCA_+_Factor_analyses.pdf
Show me some love with the like buttons below... Thank you and please don't forget to share and comment below!!
Montrez-moi un peu d'amour avec les like ci-dessous ... Merci et n'oubliez pas, s'il vous plaît, de partager et de commenter ci-dessous!
Recommended for You!
Recommended for you
This section contains best data science and self-development resources to help you on your path.
Coursera - Online Courses and Specialization
Data science
- Course: Machine Learning: Master the Fundamentals by Standford
- Specialization: Data Science by Johns Hopkins University
- Specialization: Python for Everybody by University of Michigan
- Courses: Build Skills for a Top Job in any Industry by Coursera
- Specialization: Master Machine Learning Fundamentals by University of Washington
- Specialization: Statistics with R by Duke University
- Specialization: Software Development in R by Johns Hopkins University
- Specialization: Genomic Data Science by Johns Hopkins University
Popular Courses Launched in 2020
- Google IT Automation with Python by Google
- AI for Medicine by deeplearning.ai
- Epidemiology in Public Health Practice by Johns Hopkins University
- AWS Fundamentals by Amazon Web Services
Trending Courses
- The Science of Well-Being by Yale University
- Google IT Support Professional by Google
- Python for Everybody by University of Michigan
- IBM Data Science Professional Certificate by IBM
- Business Foundations by University of Pennsylvania
- Introduction to Psychology by Yale University
- Excel Skills for Business by Macquarie University
- Psychological First Aid by Johns Hopkins University
- Graphic Design by Cal Arts
Books - Data Science
Our Books
- Practical Guide to Cluster Analysis in R by A. Kassambara (Datanovia)
- Practical Guide To Principal Component Methods in R by A. Kassambara (Datanovia)
- Machine Learning Essentials: Practical Guide in R by A. Kassambara (Datanovia)
- R Graphics Essentials for Great Data Visualization by A. Kassambara (Datanovia)
- GGPlot2 Essentials for Great Data Visualization in R by A. Kassambara (Datanovia)
- Network Analysis and Visualization in R by A. Kassambara (Datanovia)
- Practical Statistics in R for Comparing Groups: Numerical Variables by A. Kassambara (Datanovia)
- Inter-Rater Reliability Essentials: Practical Guide in R by A. Kassambara (Datanovia)
Others
- R for Data Science: Import, Tidy, Transform, Visualize, and Model Data by Hadley Wickham & Garrett Grolemund
- Hands-On Machine Learning with Scikit-Learn, Keras, and TensorFlow: Concepts, Tools, and Techniques to Build Intelligent Systems by Aurelien Géron
- Practical Statistics for Data Scientists: 50 Essential Concepts by Peter Bruce & Andrew Bruce
- Hands-On Programming with R: Write Your Own Functions And Simulations by Garrett Grolemund & Hadley Wickham
- An Introduction to Statistical Learning: with Applications in R by Gareth James et al.
- Deep Learning with R by François Chollet & J.J. Allaire
- Deep Learning with Python by François Chollet







